Inferring the Presence of Organisms
Environmental DNA (eDNA) refers to the DNA contained in living cells, in tissue fragments shed by organisms or in secretions (such as faeces and mucous) released into the environment by organisms. The analysis of eDNA in an environmental sample can be used to infer the presence of organisms in an ecosystem and has emerged as a promising tool for biodiversity assessments. For most marine organisms, eDNA analysis requires less time than traditional sampling methods such as visual surveys and is less invasive than capture approaches.
In each water sample, eDNA-containing material from multiple organisms is present. This material is filtered and then the DNA is extracted and purified. Regions of the DNA are amplified using sequence-specific barcoding primers targeting universal genetic markers. This ‘metabarcoding’ approach allows the simultaneous detection of multiple taxa in one sample. These short sections of DNA are then sequenced and compared to a reference sequence database, enabling assignment of the sequences to specific taxonomic groups.
The Metabarcoding Process
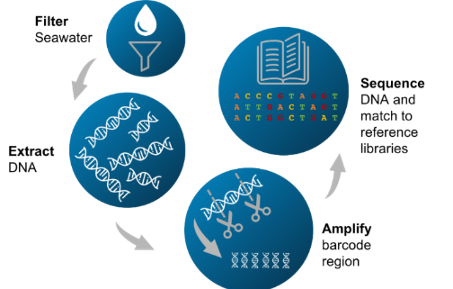
The samples from eDNA technology samples can be used for ‘omics’: to sequence and assemble environmental meta-(i.e. community) genomes, or to understand function by analysing RNA or proteins (metatranscriptomics and proteomics). Here is an overview of eDNA metbarcoding from a water sample, created by NOC's Robyn Samuel.
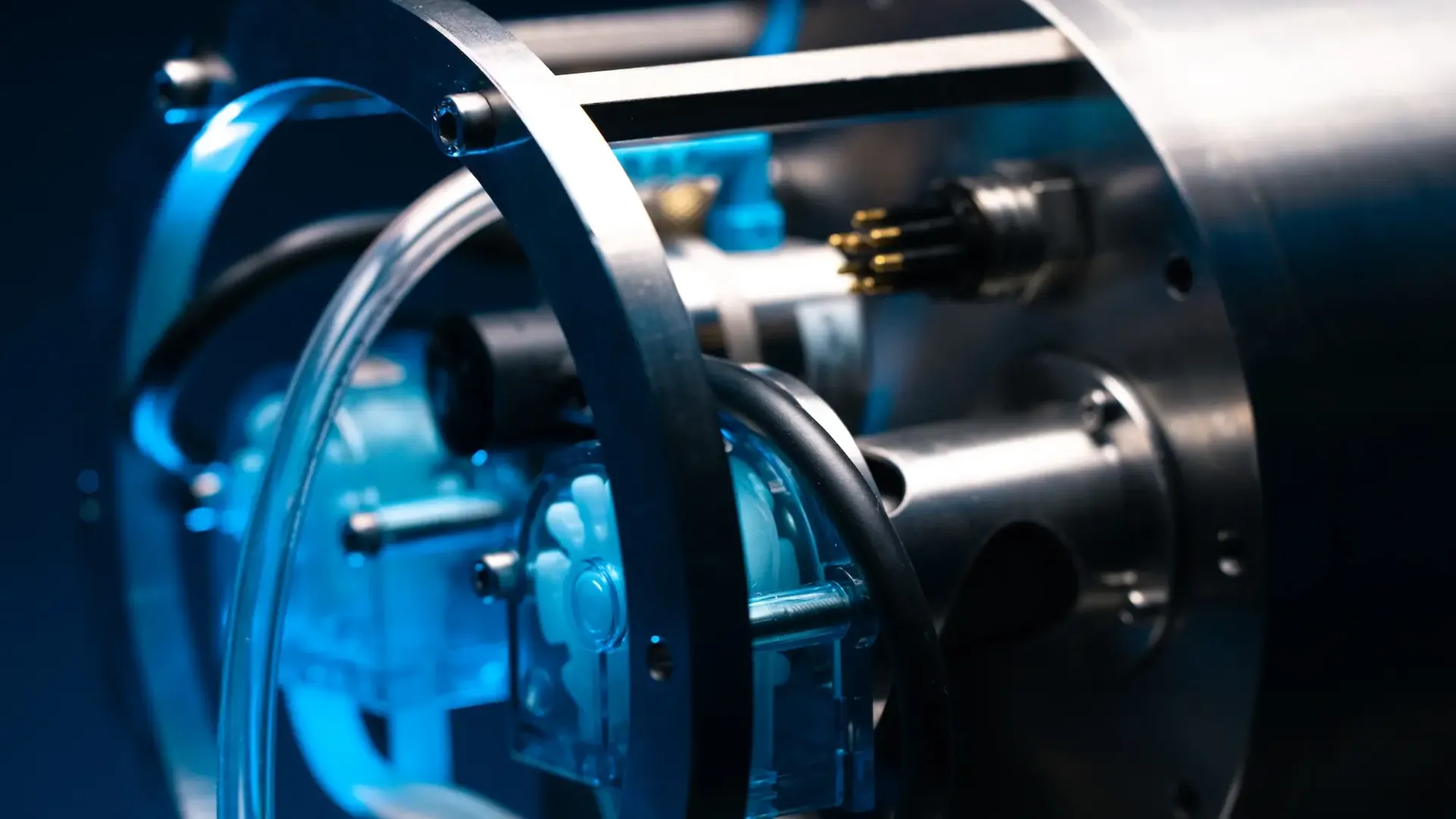
RoCSI (Robotic Cartridge Sampling Instrument)
The high-resolution eDNA sampler RoCSI, developed by the National Oceanography Centre, is an autonomous sampling device for filtering predetermined volumes of water and preserving the filters in situ. The main advantage of this is that seawater samples can be collected at high resolution and autonomously, and the filters can be immediately preserved, ensuring high quality samples are collected for eDNA metabarcoding analysis. Proof of concept deployments of RoCSI occurred in the surface ocean on the in the North Atlantic (Tang et al, 2020; Samuel et al., in prep) and in the deep sea (> 4000m depth) near Cape Verde.
More information can be found via the McLane Labs website here and here.
Current Specifications
RoCSI is capable of collecting and preserving eDNA samples from a range of biomass – from the open ocean to bloom conditions, autonomously. The rotating sample delivery mechanisms ensures that the only limitation to the number of samples one can use is the space available to store the samples. It’s rated to 6000m depth and has collected hundreds of samples across a range of platforms, including ROVs, AUVs and ASVs. A scheduling GUI allows the user to define when to sample. It has a decontamination protocol to prevent sample-to-sample carryover and pressure sensors to prevent clogging.
Functional
Depth rating: 6000m
Samples per mission: Continuous/Dependent on platform capacity
Sample types: Filtered water ≤2 L per sample, In-situ preserved Sterivex
Pore size: 0.22µm/0.45µm
Pumping system: Peristaltic pump
Decontamination system: Automated in-line bleach flushing
Weight: 15.5 kg; 10 kg wet weight
Size: 170 mm x 180 mm x 530 mm
Materials
RoCSI System: Titanium
Electronics housing: Titanium
Electrical
Power: 12V DC / 2A nominal; 16V max, 0.4 – 0.7A current at 12 V
Communication: RS-232 with cross-platform console and graphical user interfaces
Control
Operations: Set mission prior to operations; additional capacity to add decontamination runs between sampling events; sample termination based on individual thresholds (i.e. volume, time, pressure).
More Information
RoCSI Gallery
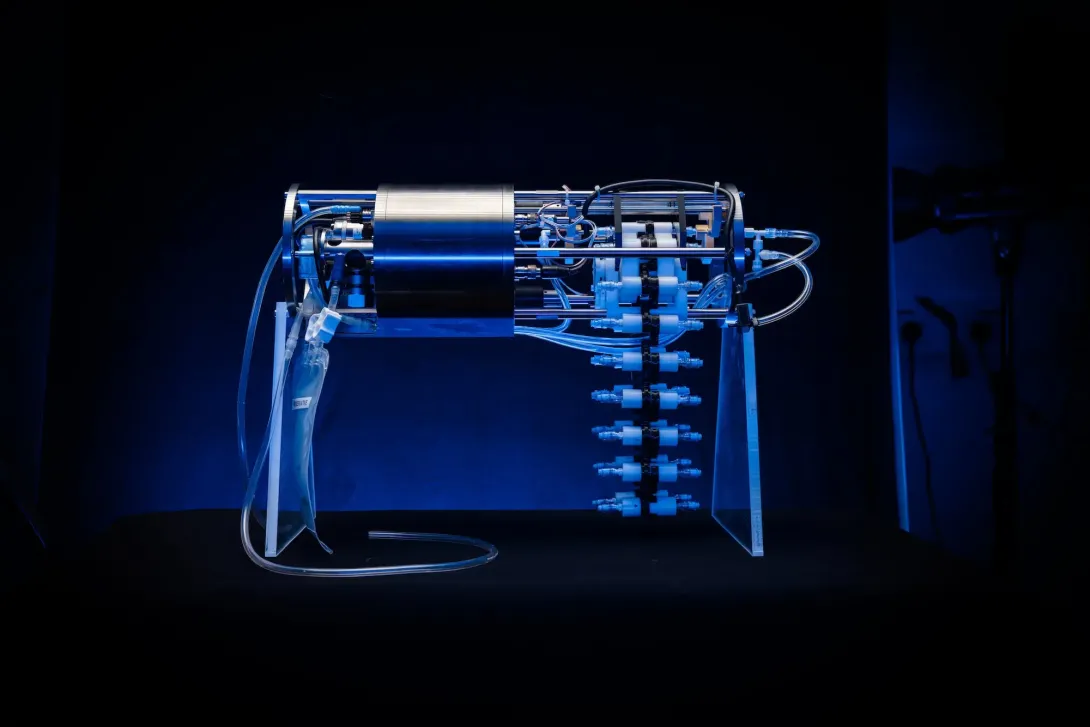
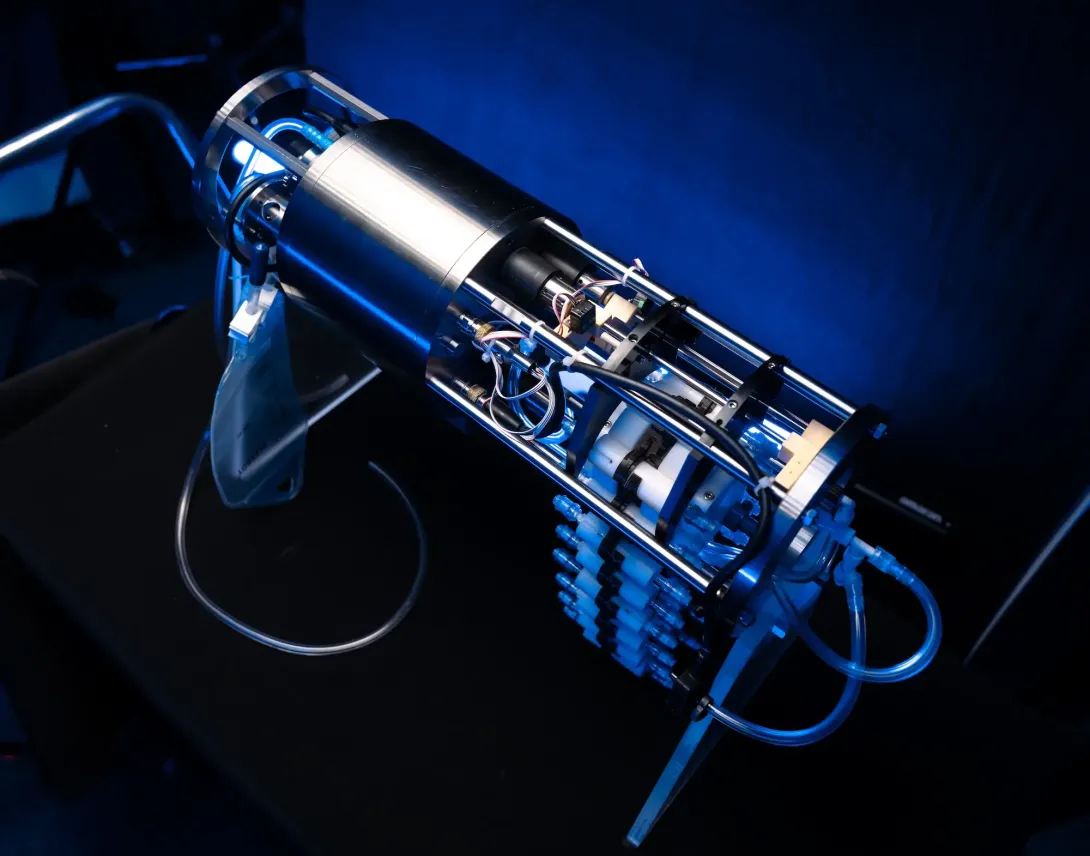
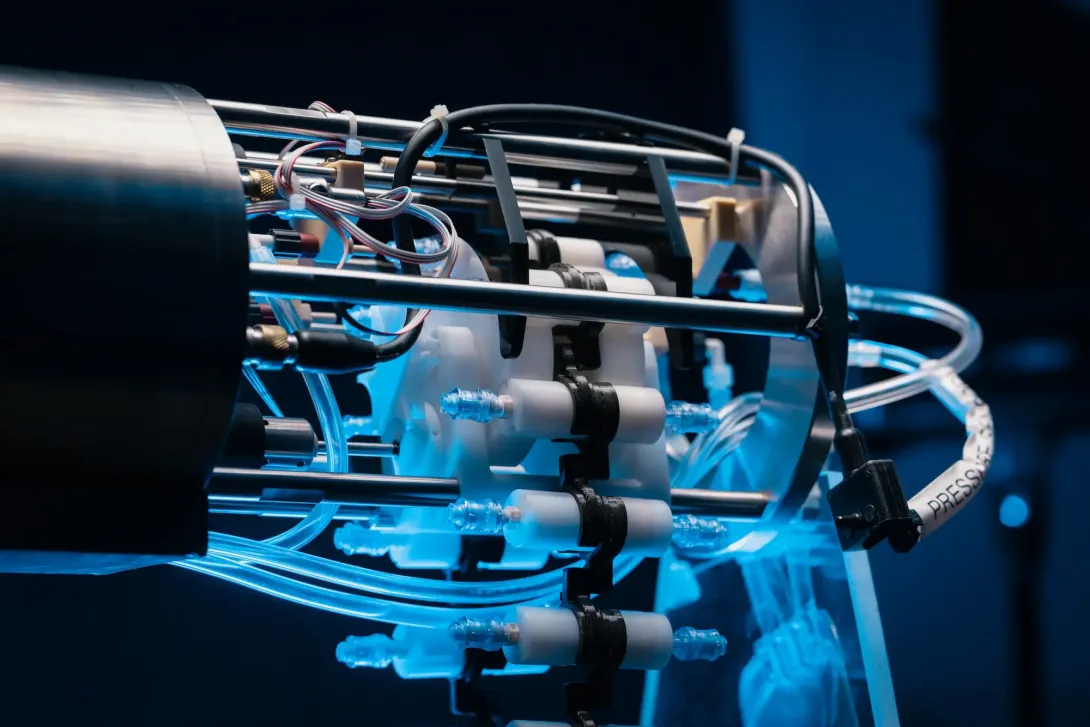
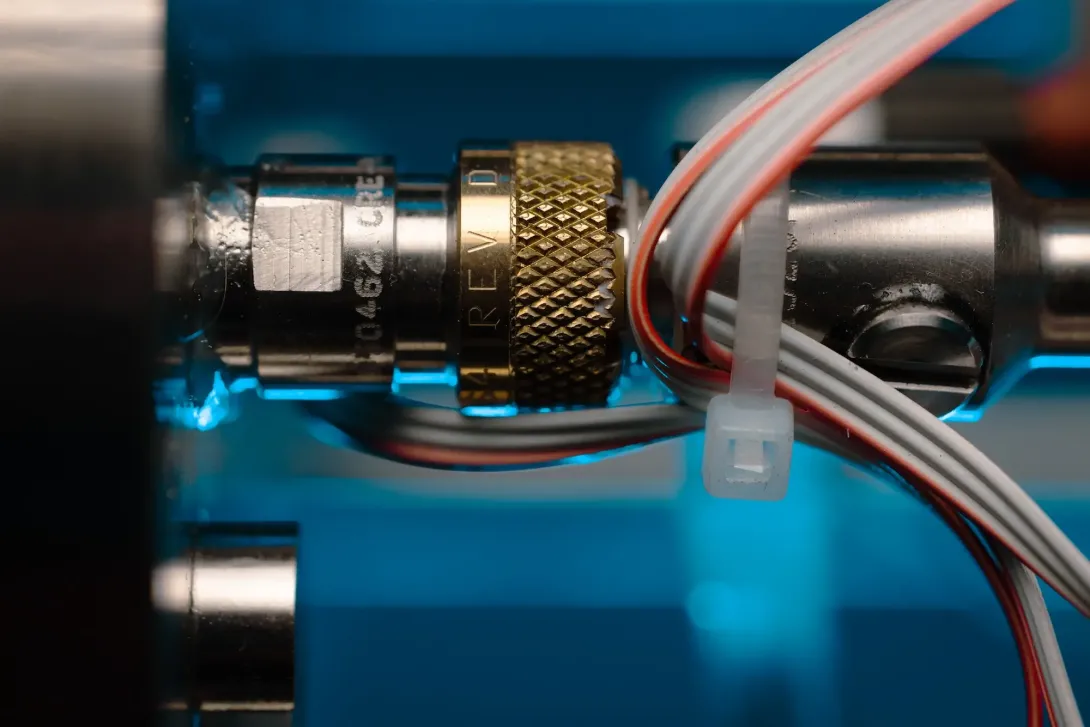




Ship's Underway eDNA Sampler
Many ships have underway systems, which pump water from just below the sea surface through the ship for analysis. We are developing an eDNA sampling system which can be integrated onto a ship’s underway system, to enable autonomous collection and preservation of eDNA from the surface water. The life here - such as phytoplankton and zooplankton - are critical for marine food webs and planetary health. This technology will let us better understand the community composition, biodiversity and function of these surface water marine ecosystems.
This new capability will add biological observations to ships of opportunity, including basin-scale longitudinal transects and UK research ships. It will save time and berths on expeditions requiring eDNA and microbiology samples, and allow us to contribute to international efforts to better monitor and understand life in the oceans.
Current Specifications
This technology filters a pre-defined volume of seawater through cartridges that capture the eDNA in the water, adds a preservative, and then labels, seals, and stores the filter. In addition to scheduled sampling (similar to RoCSI, above), this underway eDNA sampler will have adaptive sampling options as well as remote access for shore-based users to trigger additional samples based on location and environmental conditions.
Genomics Sensing In Situ
Measuring ocean biology can be an expensive and hazardous process. The oceans are vast, largely inaccessible, and normal biological measurements are dependent on the collection of samples for transit to centralised laboratories. To address these limitations, our multidisciplinary research team are developing ‘Ecogenomic’ sensor technologies, automatic analytical devices that decentralise and democratise biological data collection through reducing or removing the need for a laboratory and manual sampling. They can include deployable and submersible, or 'fieldable' (shore-based) instruments that autonomously sample, process and analyse biology directly in the environment, with near-real-time results transmitted through telemetry. This is achieved using molecular biological techniques including nucleic acid amplification-based assays for rapid (results in minutes) identification and quantification of species based on the presence of their genomic sequences within the water column. These systems remain at an early stage of development.

